The programs listed on this page are all freeware. That means that you can download them and use them without cost (as in free beer).
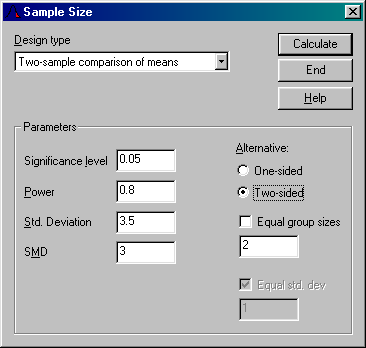
SSIZE is a small windows-program used to calculate sample sizes
for various popular (but simple) statistical designs. The programs uses a normal
distribution approximation to calculate sample sizes.
The current version of SSIZE (0.1f) can handle the following designs:
SSIZE is free and is guaranteed to do nothing.
Download the windows binaries for SSIZE v0.1f.
The rutines used in SSIZE are based on some functions
I made for S-plus a few years ago. This code has actually become the
basis for the sample-size functions in the new version of the
statistical program R.
If someone shows interest, I'll gladly include sample size calculations when looking at proportions as well as using the exact non-central t-distribution for calculating sample sizes.
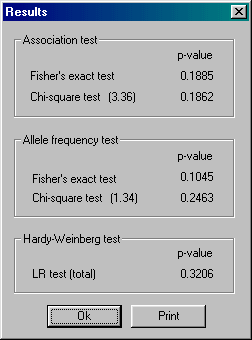
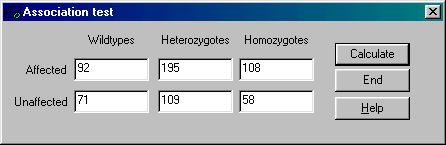 ASSOTEST is a fast windows program
to analyze association/independence, compare allele frequencies and test for
Hardy-Weinberg equilibrium for a case-control study (e.g., when a
population of affected and unaffected persons have been genotyped).
ASSOTEST is a fast windows program
to analyze association/independence, compare allele frequencies and test for
Hardy-Weinberg equilibrium for a case-control study (e.g., when a
population of affected and unaffected persons have been genotyped).
ASSOTEST can be downloaded here.
Note that you can also use Web-ASSOTEST - a web-based version of ASSOTEST, where you can also see tests for dominance, recessive, and co-dominant models. For information about assotest see the notes from the lecture about case-control studies and genetic association (PDF document).
Here's a small java applet (to the right) that can estimate the LD between two biallelic markers. All genotyped individuals are assumed independent and that there is no information about their phase.
The applet is currently being updated to include confidence limits so the version you see here may not provide the correct confidence intervals (or provide CI at all).
The printed result is R^2 (also known as Delta^2) which is one od the classical measures of linkage disequilibrium between two SNP markers. R^2 is simply the coefficient of correlation (i.e., the squared correlation coefficient) between two indices. A value of 0 corresponds to linkage equilibrium while a value close to 1 corresponds to complete linkage disequilibrium. Values close to one are a indication of complete linkage disequilibrium. A result of "NaN" indicates that insufficient data is available to estimate LD (i.e., too many cells are zero) while a red result is caused by illegal cell values (e.g., by entering letters).
Go to the PEDIPET homepage.
Go back to main page.